Drop forge_demo.mp4 or forge_demo.png into apps/landing/static/landing/img/ and it shows up here automatically.
Insilijo Science is a computational-biology practice. We build Forge — a platform for multi-omics analysis — and work with teams on specific questions where careful integration across metabolomics, microbiome, transcriptomics, proteomics, and lipidomics changes the interpretation.

Drop forge_demo.mp4 or forge_demo.png into apps/landing/static/landing/img/ and it shows up here automatically.
Forge is the Insilijo analysis environment. It takes uploaded tabular or public data, preprocesses it consistently across assays, and runs specialist modules against a unified sample-metadata model. Every analysis is reproducible — parameters captured, seeds pinned, reference snapshots SHA-verified.
Uploaded feature tables or mzML runs routed through annotation, differential, and pathway enrichment — with GIZMO as the network- level interpretation layer.
Per-feature normalisation, differential expression / abundance, pathway enrichment — sharing the same metadata and QC surface as every other assay.
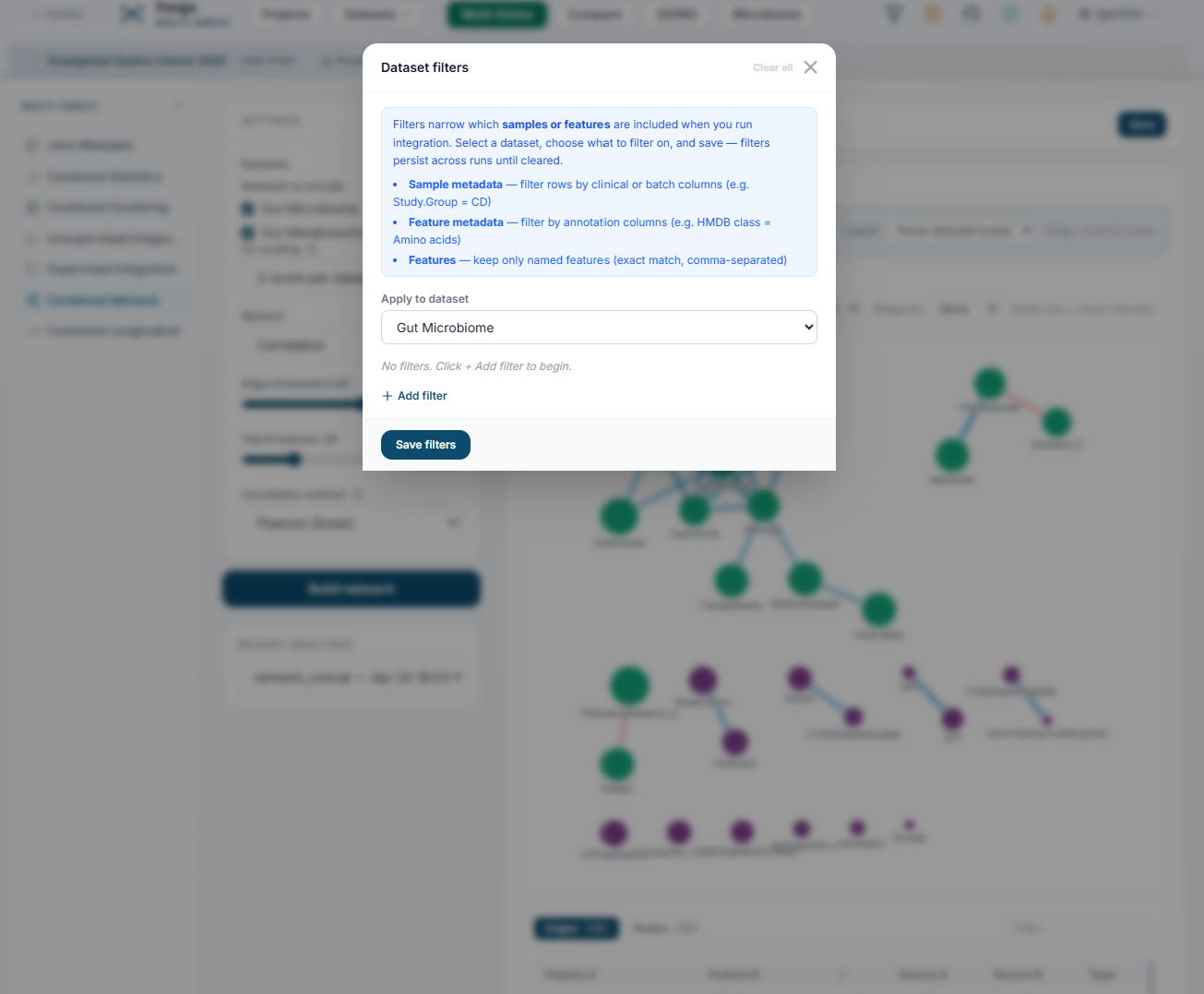
JIVE, MOFA+, SNF, MCIA, DIABLO, concatenated PCA / PLS-DA, correlation networks. Save and reload integration runs with any metadata overlay.
Lipid-class-aware preprocessing; clinical biomarker panels handled like any other tabular assay with metadata-driven stratification.
Forge is broad; these two modules are where the scientific investment has been heaviest, and where the framing advances the literature rather than just providing a familiar UI.
Community-scale integration of metagenomics and metabolomics. Reconciles the pathways encoded in a microbial community's genomes with the pathways observed in its metabolome.
A calibrated multi-omics knowledge graph spanning metabolites, reactions, genes, transcripts, proteins, phenotypes, and diseases. Takes an omics-derived signature and returns ranked diseases, causal chains, and druggable targets.
Upload your data, configure the analysis, run any Forge module. Self-serve, suitable for teams comfortable with the method.
For biotechs and research groups with a specific question. We run the analysis end-to-end and deliver interpretation, figures, and methods text ready for a manuscript or internal decision.
For PIs who want Forge modules in a paper. Co-authorship, methods writing, figure preparation. Preferred when the integration reveals something the single-assay analysis missed.
Insilijo Science is led by Joey Gardner, a computational biologist working across metabolomics, microbiome, transcriptomics, and network inference. The practice is small on purpose — most engagements run with direct principal involvement from design through manuscript.
For Joey's essays, CV, and broader reading list, see insilijo.github.io.